How deep learning extends the capabilities of cell image analysis

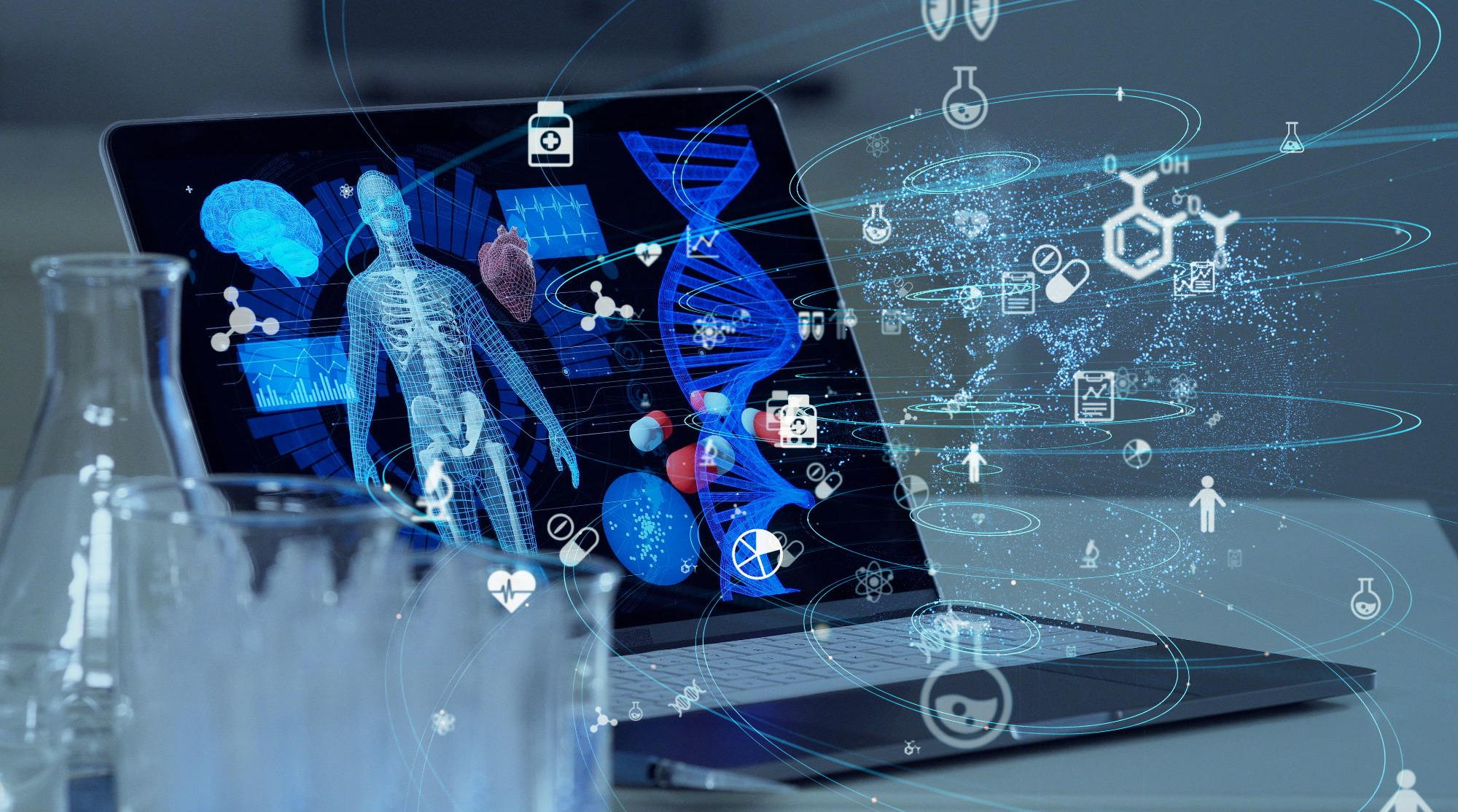
An overview of deep learning-based cell image analysis. A typical analysis pipeline consists of a retraining module and an output module: the output module directly produces the metrics. However, as the experimental setup changes, the parameters of the deep learning model must be retrained. credit: Intelligent computing (2022). DOI: 10.34133/2022/9861263
The cell is the basic structural and functional unit of life with different sizes, shapes and densities. There are many different physiological and pathological factors that influence these parameters. Therefore, it is extremely important for biomedical and pharmaceutical research to study the characteristics of cells.
Traditionally, researchers have observed cell samples directly through microscopes to study morphological changes in the cells. In recent years with the development of computer science and artificial intelligence, deep learning can now be combined with cellular analysis techniques. It can replace researchers’ direct observation under a microscope and manual interpretation of images, improving research efficiency and accuracy.
A growing number of deep learning-based algorithms have been developed to enhance the capabilities of cell image analysis, primarily to address three key challenges:
- Segmentation. To identify meaningful objects or features, the image is divided into several parts using deep learning. Cell segmentation is a fundamental prerequisite for the identification, counting, tracking, and morphological analysis of cell images;
- Tracking. That is, after the segmentation of the cell images, the behavior of the cells of the entire spectrum is monitored. Living cells contain a lot of information about living organismand dynamic characteristics of cells, especially morphological changes, can reflect the state of health of the body in pathological and physiological processessuch as immune responsewound healing, spread of cancer cells and metastases, etc.
- Classification. Classification of morphological features of cells based on extracted parameters often serves as the next analysis task for phenotypic screening and cell profiling.
A review article was published in the magazine for the above-mentioned three most important tasks Intelligent computing discusses in detail the progress of deep learning techniques.
“Unlike traditional computer vision techniques, a deep neural network (DNN) can automatically generate more effective representations than manually by learning from a large-scale data set. In cell images, deep learning-based methods also show promising results in cell segmentation and tracking,” the authors stated. “Such successful applications demonstrate the ability of DNNs to extract high-level features and shed light on the potential use of deep learning. reveal the more complex laws of life behind cellular phenotypes.”
In addition, the authors also discuss the challenges and opportunities of deep learning techniques in cell image processing. The authors stated, “Deep learning has demonstrated incredible ability to perform cell image analysis. However, there remains a significant gap in the performance of deep learning algorithms in academic research and practical applications.” Currently, there are challenges and opportunities in three aspects, namely the amount of data, data qualityand data confidence:
- Deep learning with a small but expensive data set. Generating a large-scale cell image dataset is a challenging task. This is because cell images require experienced biological experts to assign labels image by image. The scale of cell image datasets is often limited by the complexity of the annotations.
- Deep learning with noisy and unbalanced labels. The annotation quality of cell image datasets is highly dependent on the professional skills of individuals, leading to noise and label imbalance. Label noise is introduced by assigning incorrect or incomplete labels to the training images. The imbalance of labels is caused by the preference of annotations, where the number of labeled images for different classes is quite unbalanced.
- Uncertainty-Aware Cell Image Analysis. Uncertainty-aware learning is critical to the application of deep learning in biological scenarios. A conventional neural network cannot detect new phenotypes without a mechanism that reflects confidence in the classification results.
Using deep learning, scientists are exploring new technologies to improve cell image analysis. More effective solutions will be offered in the future, and deep learning and biomedical research will be more closely integrated.
Additional information:
Junde Xu et al., Deep Learning in Cell Image Analysis, Intelligent computing (2022). DOI: 10.34133/2022/9861263
Courtesy of Intelligent Computing
Citation: How Deep Learning Empowers Cell Image Analysis (2022, November 21) Retrieved November 21, 2022, from https://phys.org/news/2022-11-deep-empowers-cell-image-analysis.html
This document is subject to copyright. Except in good faith for the purpose of private study or research, no part may be reproduced without written permission. The content is provided for informational purposes only.
https://phys.org/news/2022-11-deep-empowers-cell-image-analysis.html How deep learning extends the capabilities of cell image analysis